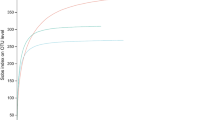Abstract
International trade in plants and climate change are two of the main factors causing damaging tree pests (i.e. fungi and insects) to spread into new areas. To mitigate these risks, a large-scale assessment of tree-associated fungi and insects is needed. We present records of endophytic fungi and insects in twigs of 17 angiosperm and gymnosperm genera, from 51 locations in 32 countries worldwide. Endophytic fungi were characterized by high-throughput sequencing of 352 samples from 145 tree species in 28 countries. Insects were reared from 227 samples of 109 tree species in 18 countries and sorted into taxonomic orders and feeding guilds. Herbivorous insects were grouped into morphospecies and were identified using molecular and morphological approaches. This dataset reveals the diversity of tree-associated taxa, as it contains 12,721 fungal Amplicon Sequence Variants and 208 herbivorous insect morphospecies, sampled across broad geographic and climatic gradients and for many tree species. This dataset will facilitate applied and fundamental studies on the distribution of fungal endophytes and insects in trees.
Measurement(s) | metagenomics analysis • Cytochrome C Oxidase Subunit 1 |
Technology Type(s) | amplicon sequencing • Dideoxy Chain Termination DNA Sequencing |
Factor Type(s) | tree species • geographic location • mean annual temperature • mean annual precipitation |
Sample Characteristic - Organism | Fungi • Insecta |
Sample Characteristic - Environment | dormant tree twigs |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.16764229
Similar content being viewed by others
Background & Summary
Fungi and insects can have large impacts on tree health, ranging from beneficial to very harmful1,2,3. In the last 200 years the number and impact of tree pests (i.e. harmful fungi and insects) has considerably increased4,5,6, mainly because non-native pests have been introduced to new areas through the global trade of plant material4,7. However, our current knowledge about the distribution of organisms associated with trees comes from studies that explored the diversity of fungi and insects at local and regional scales8,9,10,11. These studies rarely focused on more than one taxon12, or sampled multiple hosts in different geographic regions12,13,14,15. The general lack of large-scale biodiversity studies is mainly due to a lack of resources, as surveys can be laborious, costly, and often require strong collaborative networks16. Large-scale biodiversity assessments have only been done for a few organism groups, including soil fungi17, earthworms18, plants19 and terrestrial vertebrates20. Taxa associated with a single host species or genus have never been investigated in a standardized study across several continents, although this would be valuable to test general hypotheses related to global biodiversity patterns and to assess the risk associated with the global trade of plants.
We studied overwintering stages of endophytic fungi and insects associated with the twigs of multiple tree species, collected around the globe, during the winter of 2017/2018 (Fig. 1). This was done through the COST Action FP1401 “Global warning”, which aimed to create standardized protocols for the establishment and monitoring of sentinel plantings21 on a global scale, as an early warning system to detect potential tree pests. Specifically, this data set was compiled to determine which pests might be moved through trade of asymptomatic plant material. We sampled dormant twigs on multiple trees: 20 twigs were sampled per tree species and location (“a sample”). In total 145 native and non-native tree species, both angiosperms and gymnosperms, were sampled in 28 countries (total of 352 samples). Fungi were assessed by high-throughput sequencing (HTS). Insects were reared from dormant twigs collected from a subset of 109 tree species in 18 countries (total of 227 samples) and were sorted to taxonomic orders and feeding guilds. Herbivorous insects were identified by morphological and molecular methods. This data set reveals the diversity of tree-associated fungal endophytes and insects across broad geographic and climatic gradients and for many host taxa (Fig. 2). The data set can be used to investigate the biodiversity of tree-associated fungal endophytes and insects, especially in studies comparing different geographic and climatic regions and different hosts. Furthermore, the results of such analyses can be used in Pest/Pathway Risk Analyses, with the ultimate goal of reducing the likelihood of pest introductions to new areas.
Description of sample collection and sampling locations for the study. (a) Twenty twigs per tree were collected from 1–5 trees per species at each location. One sample constitutes all twigs from a given tree species at a particular location. Fungi were assessed by high-throughput sequencing from 50 mg of surface sterilized and ground tissues (i.e. pooled 0.5 cm long twig segments, halves of buds, and 0.5 cm long needle/leaf segments for evergreen species collected from a sample). Insects were reared from collected twigs and identified using morphological and molecular approaches. (b,c) Sampling locations outside Europe (b) and within Europe (c) are shown. Colors indicate the groups that were assessed from collected samples. The size of the circles is proportional to the number of samples collected at each location. The same legend applies for b and c. In several cases, samples collected at multiple locations within a country were merged for better visibility (i.e. AR, AU, EE, HU, ME, SE, TN, UK, ZA; Table 1).
Distribution of collected samples along gradients of different variables. (a) Frequency of values for pairwise comparisons of host geographic origin (geographic distance) and host species (phylogenetic distance). (b) Frequency distribution of climate variables for sites (i.e. mean annual temperature, temperature seasonality and mean annual precipitation). The size of the circles indicates the number of sample pairs (a) or samples (b). Colors indicate different groups. Geographic distance is calculated as Euclidean distance and is expressed in degrees (°) with 1° corresponding to 111 km. Phylogenetic distance is expressed in millions of years (My).
The data set includes raw sequence data obtained from HTS of the internal transcribed spacer region 2 (ITS2) of the ribosomal operon from dormant twig tissues22. We also present a sample x Amplicon Sequence Variant (ASV23) matrix, which consists of abundances of 12,721 fungal ASVs in 352 samples24 and data on the taxonomic classification of detected fungal ASVs24. We present similar data for insects: specimen counts for insects belonging to different feeding guilds (i.e. herbivores, predators and parasitoids) and taxonomic orders in each sample24 and a sample x morphospecies matrix24 showing abundances of 208 herbivorous insect morphospecies across 227 collected samples, together with data on the tentative identification of herbivorous insect morphospecies24. Aligned cytochrome c oxidase subunit 1 (COI) sequences of herbivorous insects are deposited in GenBank25. Finally, we provide a table containing information about tree species, location and climate for each sample24.
Methods
Field collection
Endophytic fungi and insects were assessed from dormant twig samples from 155 tree species at 51 locations in 32 countries. Sampled tree species belonged to genera that are native to, and occur widely across, either the northern or southern hemisphere, since very few tree genera occur naturally in both hemispheres (e.g., in our study only Podocarpus appears in both hemispheres but has a limited distribution in the northern hemisphere). We sampled largely in botanical gardens and arboreta, which allowed us to sample native and non-native, congeneric and confamiliar, tree species at each location. At each location, one native and one to three non-native congeneric or confamiliar tree species were sampled.
At each location, twenty 50-cm long asymptomatic twigs were collected from 1–5 individual trees per species, from different branches and different parts of the crown (Fig. 1). The number of individual trees per species depended on the number of trees available in the specific botanical garden or arboretum, which was often low (Table 1). All twigs per tree species and location were pooled and analysed as a single sample. On some occasions two samples of the same tree species at the same location are considered. Sampling was conducted in the month with the shortest day-length in the year (end of December 2017 in the Northern hemisphere, end of June 2018 in the Southern hemisphere). Samples originating from a tropical region (eleven samples from Tanzania) were collected in June 2018. Trees were sampled in winter to align with the timing of trade, i.e. most woody plants are traded in winter or early spring, as plants will be planted in the following spring, and to reduce the risk of introducing foliar pests in deciduous trees. Evergreen gymnosperm and angiosperm tree species, which were also considered, do not lose foliage during winter, and are thus sold with leaves/needles.
Fungal endophytes
To assess fungal communities, a total of 352 samples from 145 native and non-native tree species, belonging to nine families of angiosperms and gymnosperms, were collected. Sampling was done at 44 locations in 28 countries on five continents (Fig. 1, Table 1).
From each twig in a sample, one bud, one needle/leaf and one 1 cm long twig segment were taken (Fig. 1). Needles from gymnosperms, and leaves from evergreen angiosperms were sampled to accurately assess the risk of trading these species. Twig segments were cut from the twig bases. The selected plant parts were surface sterilized by immersion in 75% ethanol for 1 min, 4% NaOCl for 5 min, and 75% ethanol for 30 s26. After air drying on a sterile bench, the following material from each of 20 twigs per sample was pooled: half of one bud, a 0.5 cm long piece of a needle (from gymnosperms) or a 0.25 cm2 leaf (for evergreen angiosperms) and a 0.5 cm long piece of twig.
DNA extraction, PCR amplification and Illumina sequencing
Total genomic DNA was extracted from 50 mg of pooled, surface sterilized, and ground tissue (Fig. 1) using DNeasy PowerPlant Pro Kit (Qiagen, Hilden, Germany), following the manufacturer’s instructions. For a total of 31 out of 352 samples, DNA was extracted from different tissues separately, and DNA extracts were then pooled. DNA concentrations were quantified using the Qubit dsDNA BR Assay Kit (Thermo Fisher Scientific, Waltham, USA) on a Qubit 3.0 Fluorometer (Thermo Fisher Scientific) and DNA was diluted to 5 ng/μl. Samples that yielded less than 5 ng/μl were not diluted. The ITS2 region was amplified with the 5.8S-Fung and ITS4-Fung primers27. PCR amplifications were carried out in 20 μl reaction volumes containing 25 ng of DNA template, 1 mg/ml BSA, 1 mM of MgCl2, 0.4 μM of each primer, and 0.76 × JumpStart REDTaq ReadyMix Reaction Mix (Sigma-Aldrich, Steinheim, Germany). PCR was performed using Veriti 96-Well Thermal Cycler (Applied Biosystems, Foster City, CA, USA) as described in Franić et al. (2019). Each sample was amplified in triplicates and successful PCR amplification confirmed by visualization of the PCR products, before and after pooling the triplicates, on 1.5% (w/v) agarose gel with ethidium bromide staining. Pooled amplicons were sent to the Génome Québec Innovation Center at McGill University (Montréal, Quebec, Canada) for barcoding using Fluidigm Access Array technology (Fluidigm, South San Francisco, CA, USA) and paired-end sequencing on the Illumina MiSeq v3 platform (Illumina Inc., San Diego, CA, USA). Raw sequences obtained in this study are deposited at the NCBI Sequence Read Archive under BioProject accession number PRJNA70814822.
Bioinformatics and taxonomical classification of ASVs
Quality filtering and delineation into ASVs were done with a customized pipeline28 largely based on VSEARCH29, as described by Herzog et al.30. The output data available on Figshare show the abundances of fungal ASVs in the samples24. Taxonomic classification of ASVs was conducted using Sintax31 implemented in VSEARCH against the UNITE v.7.2 database32 with a bootstrap support of 80%. The data on the taxonomic classification of fungal ASVs is deposited in Figshare24.
Quality filtering, delineation into ASVs, and taxonomical assignments were done on a larger data set (total of 474 samples), which increased the confidence in the selected centroid sequences. This data set consisted of (1) sequences obtained from 352 samples of pooled tree tissues that are presented here22, (2) sequences obtained from 33 samples of pooled tree tissues which were not included in this manuscript due to violation of the common protocol, (3) sequences from 21 contaminated samples (positive DNA extraction controls), including sequences from the two control samples (not presented here), and (4) sequences obtained from 66 samples of non-pooled tree tissues of Pinus sylvestris and Quercus robur that were collected from the subset of locations considered in this study, but for a different study, and are thus not presented here.
Herbivorous insects
Insects were assessed from 227 samples of 109 tree species, collected at 31 locations and in 18 countries (Fig. 1, Table 1).
The collected twigs (twenty 50 cm twigs per species per location) were brought to a laboratory close to each sampling location and inspected for the presence of insects that overwinter as adults. Twigs were kept at room temperature with the cut ends immersed in water to induce budding and to allow the development of insects that overwinter as larvae, pupae or eggs. Twigs from each sample were protected with gauze bags to prevent insects moving between samples (Fig. 1). Twigs were inspected for the presence of insects daily for 4 weeks and all collected insects were stored in 95% ethanol for further examination.
Morphological and molecular identification
Insects were inspected using a stereo microscope and sorted to taxonomic orders and feeding guilds (i.e. herbivores, predators, parasitoids and other). The abundance of the different feeding guilds and taxonomic orders in the samples is presented in a file deposited on Figshare24. Herbivorous insects were further sorted into morphospecies and at least one specimen per morphospecies was stored at −20 °C for molecular analysis. The abundance of the different morphospecies in each sample is presented in a file deposited on Figshare24. Specimens for molecular analysis were photographed with a Leica DVM6 digital microscope and the Leica Application Suite X (LAS X). Depending on the size of the insects, the whole individual or parts (e.g. legs, head) were used for molecular analysis. Genomic DNA was extracted with a KingFisher (Thermo Fisher Scientific) extraction protocol suitable for insects (35 min incubation at RT, 30 min wash at RT with 3 different washing buffers, 13 min elution at 60 °C) in a 96-well plate. PCR for the COI was carried out in 25 µl reaction volume with 2 µl diluted DNA (1:10), 0.5 µM of each of the primers LCO1490 and HCO219833 and 1 x REDTaq ReadyMix Reaction Mix (Sigma-Aldrich) using a Veriti 96-Well Thermal Cycler (Applied Biosystems) with the following setting: 2 min at 94 °C, five cycles of 30 s at 94 °C, 40 s at 45 °C, and 1 min at 72 °C, 35 cycles of 30 s at 94 °C, 50 s at 51 °C, and 1 min at 72 °C, and a final extension step at 72 °C for 10 min. The success of amplification was verified by electrophoresis of the PCR products in 1.5% (w/v) agarose gel at 90 V for 30 min with ethidium bromide staining. A standard Sanger sequencing of the PCR products in both directions with the same primers was done at Macrogen Europe, Amsterdam, Netherlands. Sequences were assembled and edited with CLC Workbench (Version 7.6.2, Quiagen) and compared to reference sequences in BOLD34. If no conclusive results were found, sequences were compared to reference sequences in the National Centre for Biotechnology Information (NCBI) GenBank databases35. Specimens were assigned to species if the query sequence showed less than 1% divergence from the reference sequence. If two or more taxa matched within the same range, the assignment was ranked down to the next taxonomic level (i.e., genus). When no species match was obtained based on the above criteria, a genus was assigned with a divergence of less than 3%. For lower taxonomic groups the 100 nearest sequences were inspected on the Blast Tree (Fast Minimum Evolution Method) and the taxonomic relationship was evaluated based on that tree. If none of the approaches above revealed a conclusive taxonomic assignment, the morphological identification was taken as reference. The results of morphological and molecular identification of insect specimens are presented in a file deposited on Figshare24. Insect sequences are deposited in GenBank database under accession numbers MW441337-MW44176725.
Sample metadata
Pairwise geographic distances (Euclidean distances) between sampling locations were calculated based on the geographic coordinates of the locations, with function “dist” in the R statistical programme36.
Climate data, including mean annual temperature, mean annual precipitation, and temperature seasonality were obtained from the WorldClim database37, at a resolution of 2.5 min, and represent averages between 1970 and 2000.
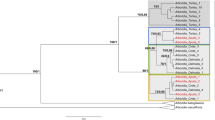
A host-tree phylogeny was constructed with the phylomatic function from the package brranching38 in R using the “zanne2014” reference tree39. One Eucalyptus sample collected in Argentina and two Eucalyptus samples collected in Tunisia were not identified to species. To place them in the phylogeny, we assigned them to different congeneric species that were not sampled in this study and that we considered as representative samples of phylogenetic diversity from across Eucalyptus genus (E. viminalis, E. robusta and E. radiata). Pairwise phylogenetic distances between study tree species were calculated using the “cophenetic” function in R36.
The described sample metadata are available in a file on Figshare24.
Data Records
The raw paired-end Illumina sequencing reads of the ITS2 region obtained from fungal DNA extracted from pooled twig tissues are archived at the NCBI Sequence Read Archive under BioProject accession number PRJNA70814822. For each of 352 samples, two fastaq files are deposited, corresponding to reverse and forward sequence reads.
The excel file “Sample x fungal Amplicon Sequence Variant matrix” shows the distribution of 11,613,187 sequence reads of 12,721 fungal ASVs among 352 samples that were used for the fungal assessment and is available on Figshare24.
An Excel table with the identity of obtained fungal ASVs (i.e., “Identity of fungal Amplicon Sequence Variants”) is also available on Figshare24. The ASVs belonged to six phyla, with the majority belonging to the Ascomycota (68%) and the Basidiomycota (15%). Around 17% of ASVs could not be assigned to a phylum. Furthermore, 9,150 ASVs (72%) were assigned to one of 28 identified classes. The five most numerous classes were: Dothideomycetes (26%), Sordariomycetes (9%), Eurotiomycetes (9%), Leotiomycetes (7%), Tremellomycetes (6%). Only 31% of the ASVs were assigned to one of 661 identified genera, and 14% to one of 760 species.
A total of 4,751 insect specimens were reared from 227 samples. The number of insects in different feeding guilds and orders is provided for each sample from which insects were collected (i.e. 154 out of 227 samples) in the excel file “Insect specimens per feeding guild, taxonomic order and sample” and is stored on Figshare24. Almost 64% of collected specimens were classified as herbivores (3,032), around 10% were classified as parasitoids (499) and only 2% were predators (89). An additional 1,131 out of 4,751 (24%) specimens could not be assigned to a feeding guild.
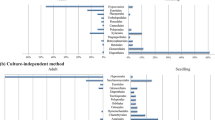
An excel file showing abundances of 3,032 specimens of herbivorous insects, belonging to 208 morphospecies, in 227 collected samples (i.e., “Sample x herbivorous insect morphospecies matrix”) is provided on Figshare24. Herbivorous insects were from seven orders: Hemiptera (78.0%), Hymenoptera (11.9%), Lepidoptera (7.1%), Coleoptera (1.3%), Diptera (0.9%), Thysanoptera (0.7%) and Orthoptera (0.1%). A table containing data on the tentative identification of representative specimens of herbivorous insect morphospecies24 is also provided. A total of 554 specimens were sequenced and 431 yielded good quality sequences. Of those, 206 sequences were assigned to species, and an additional 123 were assigned to a genus. Assembled herbivorous insect COI sequences are deposited in GenBank database under accession numbers MW441337-MW44176725.
Data records on host identity (i.e. host species, hemisphere of origin and native or non-native host distribution range), geographic location (i.e. country of collection, location, latitude and longitude) and climate (i.e. mean annual temperature, temperature seasonality, mean annual precipitation) are also provided for each sample in the excel file “Sample Metadata” on Figshare24.
Technical Validation
Sampling and sample processing were done in a standardized way as described above. All samples that were not collected and processed following the protocols were discarded (33 fungal samples). Negative controls were used for DNA extractions and PCRs to ensure the absence of external and cross contamination, and if controls were positive, samples were not used (21 fungal sample). Positive PCR controls were used to ensure that PCRs were performed correctly.
Code availability
R functions and databases used for generating the sample metadata are specified in the method section. A customized pipeline used for quality filtering of the raw sequence data obtained from HTS, delineation into ASVs and taxonomic classification of ASVs as described in Herzog et al.30 is available as a “ITS2.bash” file from the Zenodo repository28.
References
Rodriguez, R. J., White, J. F. J., Arnold, A. E. & Redman, R. S. Fungal endophytes: diversity and functional roles. New Phytol. 182, 314–330 (2009).
Dajoz, R. Insects and forests: the role and diversity of insects in the forest environment. (Intercept, 2000).
Stone, J. K., Polishook, J. D. & White, J. F. J. Endophytic fungi. in Biodiversity of fungi: Inventory and Monitoring Methods (eds. Mueller, G., Bills, G. & Foster, M.) 241–270, https://doi.org/10.1016/B978-0-12-509551-8.50015-5 (Elsevier Inc., 2004).
Santini, A. et al. Biogeographical patterns and determinants of invasion by forest pathogens in Europe. New Phytol. 197, 238–250 (2013).
Eschen, R., Roques, A. & Santini, A. Taxonomic dissimilarity in patterns of interception and establishment of alien arthropods, nematodes and pathogens affecting woody plants in Europe. Divers. Distrib. 21, 36–45 (2015).
Boyd, I. L., Freer-Smith, P. H., Gilligan, C. A. & Godfray, H. C. J. The consequence of tree pests and diseases for ecosystem services. Science. 342, 1235773 (2013).
Desprez-Loustau, M.-L. Alien Fungi of Europe. in DAISIE Handbook of Alien Species in Europe (ed. Drake, J. A.) 15–29 (Springer International Publishing, 2009).
Novotny, V. et al. Low beta diversity of herbivorous insects in tropical forests. Nature 448, 692–695 (2007).
Nguyen, D. et al. Fungal disease incidence along tree diversity gradients depends on latitude in European forests. Ecol. Evol. 6, 2426–2438 (2016).
Nguyen, D. et al. Foliar fungi of Betula pendula: Impact of tree species mixtures and assessment methods. Sci. Rep. 7, 1–11 (2017).
Vincent, J. B., Weiblen, G. D. & May, G. Host associations and beta diversity of fungal endophyte communities in New Guinea rainforest trees. Mol. Ecol. 25, 825–841 (2016).
Franić, I. et al. Are traded forest tree seeds a potential source of nonnative pests? Ecol. Appl. 29, e01971 (2019).
Arnold, A. E. & Lutzoni, F. Diversity and host range of foliar fungal endophytes: Are tropical leaves biodiversity hotspots? Ecology 88, 541–549 (2007).
U’Ren, J. M. et al. Host availability drives distributions of fungal endophytes in the imperilled boreal realm. Nat. Ecol. Evol. 3, 1430–1437 (2019).
Cleary, M., Oskay, F., Doğmuş, H. T., Woodward, S. & Vettraino, A. M. Cryptic risks to forest biosecurity associated with the global movement of commercial seed. Forests 10, 459 (2019).
Eschen, R. et al. Spotting the pests of tomorrow—Sampling designs for detection of species associations with woody plants. J. Biogeogr. 46, 2159–2173 (2019).
Tedersoo, L. et al. Global diversity and geography of soil fungi. Science. 346, 6213 (2014).
Phillips, H. R. P. et al. Global distribution of earthworm diversity. Science. 366, 480–485 (2019).
Kier, G. et al. Global patterns of plant diversity and floristic knowledge. J. Biogeogr. 32, 1107–1116 (2005).
Jenkins, C. N., Pimm, S. L. & Joppa, L. N. Global patterns of terrestrial vertebrate diversity and conservation. Proc. Natl. Acad. Sci. USA 110, E2603–E2610 (2013).
Eschen, R. et al. Safeguarding global plant health: the rise of sentinels. J. Pest Sci. (2004). 92, 29–36 (2019).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP309894 (2021).
Callahan, B. J., McMurdie, P. J. & Holmes, S. P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 11, 2639–2643 (2017).
Franić, I. et al. Worldwide diversity of endophytic fungi and insects associated with dormant tree twigs. figshare https://doi.org/10.6084/m9.figshare.c.5353304 (2022).
Genbank https://identifiers.org/ncbi/insdc:MW441337 (2021).
Toti, L., Viret, O., Horat, G. & Petrini, O. Detection of the endophyte Discula umbrinella in buds and twigs of Fagus sylvatica. Eur. J. For. Pathol. 23, 147–152 (1993).
Taylor, D. L. et al. Accurate estimation of fungal diversity and abundance through improved lineage-specific primers optimized for illumina amplicon sequencing. Appl. Environ. Microbiol. 82, 7217–7226 (2016).
Hartmann, M. martin-hartmann/metabarcoding: metabarcoding_20211019. Zenodo https://doi.org/10.5281/zenodo.5578643 (2021).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 10, 1–22 (2016).
Herzog, C. et al. Microbial succession on decomposing root litter in a drought-prone Scots pine forest. ISME J. 13, 2346–2362 (2019).
Edgar, R. SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv 074161 https://doi.org/10.1101/074161 (2016).
Abarenkov, K. et al. The UNITE database for molecular identification of fungi – recent updates and future perspectives Rationale. New Phytol. 186, 281–285 (2010).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3, 294–9 (1994).
Ratnasingham, S. & Hebert, P. D. N. BOLD: The Barcode of Life Data System. Mol. Ecol. Notes 7, 355–364, http://www.barcodinglife.org (2007).
Geer, L. et al. The NCBI BioSystems database. Nucleic Acids Res. 38 (2010).
R Core Team. R: A language and environment for statistical computing. (R Foundation for Statistical Computing, 2018).
Fick, S. E. & Hijmans, R. J. Worldclim 2: New 1-km spatial resolution climate surfaces for global land areas. Int. J. Climatol. 37, 4302–4315 (2017).
Chamberlain, S. brranching: Fetch ‘Phylogenies’ from Many Sources. R Packag. version 0.5.0. (2019).
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing environments. Nature 506, 89–92 (2014).
Acknowledgements
We gratefully acknowledge the financial support of the Swiss National Science Foundation (Project C15.0081) grant number 174644 and the Swiss Federal Office for the Environment (Grant 00.0418.PZ/ P193-1077). This work was supported by COST Action “Global Warning” (FP1401). CABI is an international intergovernmental organisation, and R.E., M.K., H.L. and I.F. gratefully acknowledge the core financial support from our member countries (and lead agencies) including the United Kingdom (Foreign, Commonwealth & Development Office), China (Chinese Ministry of Agriculture and Rural Affairs), Australia (Australian Centre for International Agricultural Research), Canada (Agriculture and Agri-Food Canada), Netherlands (Directorate General for International Cooperation), and Switzerland (Swiss Agency for Development and Cooperation). See https://www.cabi.org/aboutcabi/who-we-work-with/key-donors/ for full details. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. M.B. and M.K.H. were financially supported by the Slovak Research and Development Agency (Project APVV-19-0116). H.B. would like to thank the botanist Jorge Capelo who helped with Myrtaceae identification and INIAV IP for supporting her contribution to this study. Contributions of M. de G. and B.P. were financed through Slovenian Research Agency (P4-0107) and by the Slovenian Ministry of Agriculture, Forestry and Food (Public Forestry Service). G.C, C.B.E. and A.F.M. were supported by OTKA 128008 research grant provided by the National Research, Development and Innovation Office. Contributions of K.A. and R.D. were supported by the Estonian Science Foundation grant PSG136. M.J.J., C.L.M. and H.P.R. were financially supported by the 15. Juni Fonden (Grant 2017-N-123). P.B., B.G. and M.Ka. were financially supported by the Ministry of Science and Higher Education of the Republic of Poland for the University of Agriculture in Krakow (SUB/040013-D019). C.N. was financially supported by the Slovak Research and Development Agency (Grant APVV-15-0531). N.K. was partially supported by Russian Foundation for Basic Research (Grants 15-29-02645, 19-04-01029). R.OH. was supported by funding from DAERA, and assistance from David Craig, AFBI. T.P. thanks the South African Department of Forestry, Fisheries and the Environment (DFFE) for funding noting that this publication does not necessarily represent the views or opinions of DFFE or its employees. In preparing the publication, materials of the bioresource scientific collection of the CSBG SB RAS “Collections of living plants indoors and outdoors” USU_440534 (Novosibirsk, Russia) were used. M.Z. was financially supported by Ministry of education, science and technological development of the Republic of Serbia (Contract 451-03-68/2020-14/200197). We acknowledge the Genetic Diversity Centre (GDC) at ETH Zurich for providing computational infrastructure and acknowledge the contribution of McGill University and Génome Québec Innovation Center (Montréal, Quebec, Canada) for pair-end sequencing on Illumina MiSeq.
Author information
Authors and Affiliations
Contributions
I.F., S.P. and R.E. designed the study. I.F. compiled the common sampling and laboratory protocols and coordinated sample collection. All co-authors did sample collection and/or DNA extractions for fungal assessment and/or insect rearing. I.F finalized laboratory preparation of samples for HTS. M.H. supervised amplicon sequencing approach and performed the bioinformatic analyses of the HTS sequence data. N.F. and B.R. did morphological and molecular identification of insect with support of M.M.G.. I.F., S.P. and R.E. wrote the manuscript with contribution from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Franić, I., Prospero, S., Adamson, K. et al. Worldwide diversity of endophytic fungi and insects associated with dormant tree twigs. Sci Data 9, 62 (2022). https://doi.org/10.1038/s41597-022-01162-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-022-01162-3
This article is cited by
-
Climate, host and geography shape insect and fungal communities of trees
Scientific Reports (2023)